IIT Bombay and IIT Mandi researchers develop robotic model to decode microbial swimming behaviour
Tabletop Robot Twins Reveal How Microbes Perform Run-and-Tumble Motion
Mumbai | January 2026
In a groundbreaking study inspired by biomimicry, researchers from the Indian Institute of Technology (IIT) Bombay and IIT Mandi have successfully developed a tabletop robotic system that accurately reproduces the run-and-tumble motion—a fundamental navigation pattern used by microbes such as bacteria and algae.
The study, published in the prestigious journal Physical Review Letters, provides new insights into microbial motility, one of the most basic yet mysterious processes in living systems.
Understanding Run-and-Tumble Motion
Run-and-tumble motion is commonly observed in microorganisms like Chlamydomonas reinhardtii, a single-celled alga. These organisms swim in a straight line (run) and suddenly change direction (tumble), allowing them to navigate complex environments.
According to Prof. Nitin Kumar (IIT Bombay), Chlamydomonas swims using two flagella that usually beat in sync. When this synchronization breaks, the organism abruptly changes direction, resulting in a tumble.
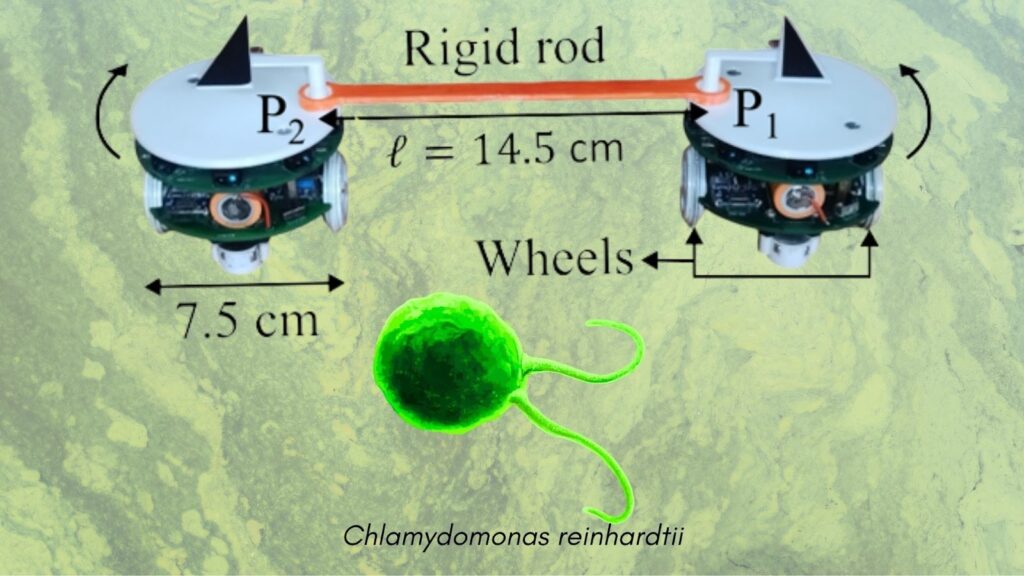
Robots Mimic Microbial Flagella
To replicate this behaviour, the research team replaced biological flagella with two self-propelled robots mechanically coupled by a rigid rod—an analogue of the distal fibre that connects flagella in real microorganisms.
Lead authors Somnath Paramanick (IIT Bombay) and Umashankar Pardhi (IIT Mandi) designed the system under the guidance of Prof. Nitin Kumar and Prof. Harsh Soni (IIT Mandi).
By adjusting:
- the angle of connection, and
- the distance of the pivot point from the robot’s centre,
the researchers were able to control tumbling frequency and running stability, closely mirroring microbial motion.
Simulating the Microbial World
Microorganisms live in a world dominated by friction rather than inertia. To recreate this environment, the robots were placed on a high-friction surface, simulating overdamped active Brownian motion, where momentum is negligible.
“This setup allows macroscale robots to obey the same physical laws as microscopic swimmers,” explains Prof. Kumar.
Key Findings
- Robots exhibited long straight runs interrupted by sharp tumbles, often with near 180-degree direction reversal
- Run durations closely matched those observed in real bacteria and algae
- Tumbling emerged spontaneously, without external programming
- Hydrodynamic interactions were not required, challenging previous assumptions in microbiology
Scientific and Practical Impact
The study demonstrates that complex biological motion can emerge from simple physical interactions, offering a new framework for studying active matter systems.
“This approach helps explain how different swimming modes arise purely from mechanical coupling,” notes Prof. Harsh Soni.
The findings could also contribute to the development of autonomous microscopic robots for medical diagnostics, drug delivery, and healthcare applications.
Implications for Biology
The researchers suggest that real microorganisms may actively regulate the mechanical properties of their internal fibres to switch between running and tumbling modes—an insight that could reshape our understanding of microbial behaviour.




